 |
|
|

|
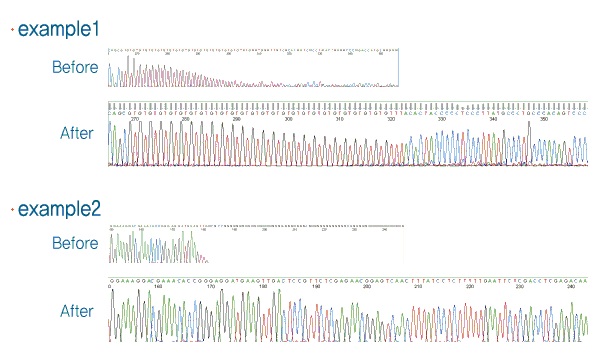
Psomagen offers DNA sequencing for difficult templates with high GC or GT, unusual secondary structure or hairpin structures. With over 20 years of experiences, we have optimized our protocol to yield successful sequencing data for these difficult templates.
|
| Psomagen provides premium protocol for difficult template sequencing.
|
- High GC or GT
- Unusual secondary structure
- Repetitive sequence
- siRNA with hairpin structure
- Homopolymeric tracts (PolyA, PolyG)
|
|
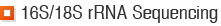
|
 |
Psomagen offers sequencing service for 16S/18S rRNA gene from your bacterial or fungal colonies rapidly and effectively. We provide all-inclusive service performing DNA extraction, PCR, sequencing and assembly.
|
|
 |
|

|
Performs all processes including nucleic acid extraction, PCR,
sequencing, assembly using bacterial cells. |
 |
Deliver
results within 7 business days from date
of sample arrival |
|
| |
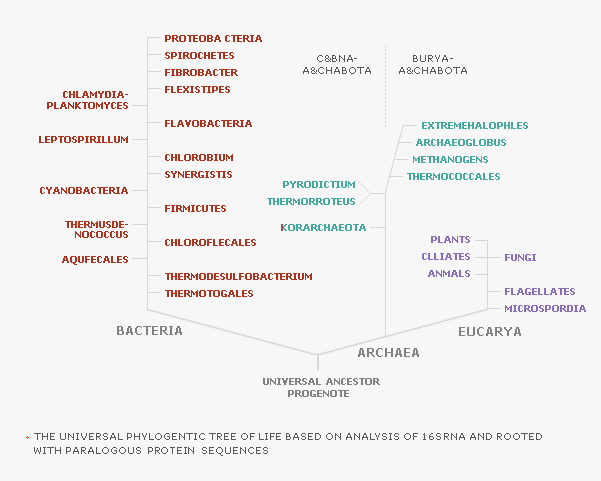
| As shown in the figure below, living organisms
can be classified into 3 primary lineages using the base sequence
analysis of 16S rRNA: Domain Eucarya, Domain Bacteria, and Domain
Archaea. |
 |
|
 |
 |
| Psomagen offers primer walking service to confirm the sequence integrity of your clone or PCR products. We can perform end sequencing with the primers that you provide or use the reference sequence, if applicable, and design internal primers to fill in the gap. |
 |
|
 |
|
 |
1Kb single contig : 5~6 days. |
| |
(Time consuming would vary based on the following
reasons; primer synthesis and sequencing |
| |
failure) |
 |
Phred score of 20 or higher quality contig is
requirerd. |
 |
Time can be saved when primers are synthesized
bidirectionally. |
| |
(refer to the figure as
follows) |
|
| |
 |
| |
 |
Starting with the end sequencing product by using the primer provided or designed, |
| |
internal primer will be designed and processed. |
| |
|
 |
Using a new primer (but with the same template), internal primer
continues reading at |
| |
the appropriate location. |
| |
|
 |
Each walking requires approximately 5~6
days; enabling extension of 500bp~600bp per direction. |
|
| |
 |
|
|
|
|
